Orientation¶
Loading and Saving Data¶
File Formats¶
This software package works with NIFTI volumes. Some builds may contain experimental support for folders of DICOM files, however this is not well tested.
Alternative packages which are able to convert DICOM files to NIFTI include the following:
- itk-snap
- dcm2nii
- Or the batch version which allows a number of volumes to be converted dcm2niibatch
Loading data using Drag and Drop¶
You can drag and drop single or multiple files onto the main window to load data. You will be prompted to choose the type of data:
The suggested name is derived from the file name but is modified to ensure that it is a valid name (data names must be valid Python variable names) and does not clash with any existing data.
If you choose a name which is the same as an existing data set, you will be asked if you wish to overwrite the existing data.
When dropping multiple files you will be asked to choose the type of each one. If you select cancel the data file will not be loaded.
Saving Data¶
The following menu options are used for saving data:
- File -> Save current data
- File -> Save current ROI
So, to save a data set you need to make it the current data, using the Overlay menu or the Volumes widget. Similarly to save an ROI you need to make it the current ROI. Saving the main data can be done by selecting it as the current overlay.
Save a screen shot or plot¶
- Right click on an image or plot
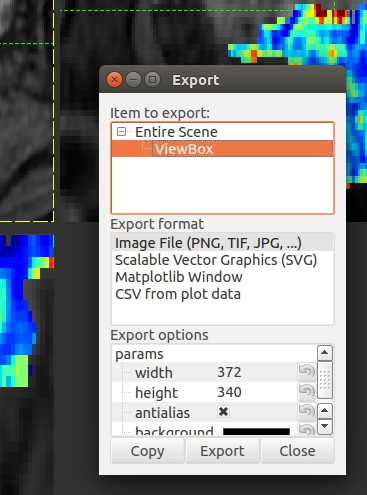
- Click Export
- A view box will appear with the various format options.
- svg format will allow editing of the layers and nodes in inkscape or another vector graphics viewer.
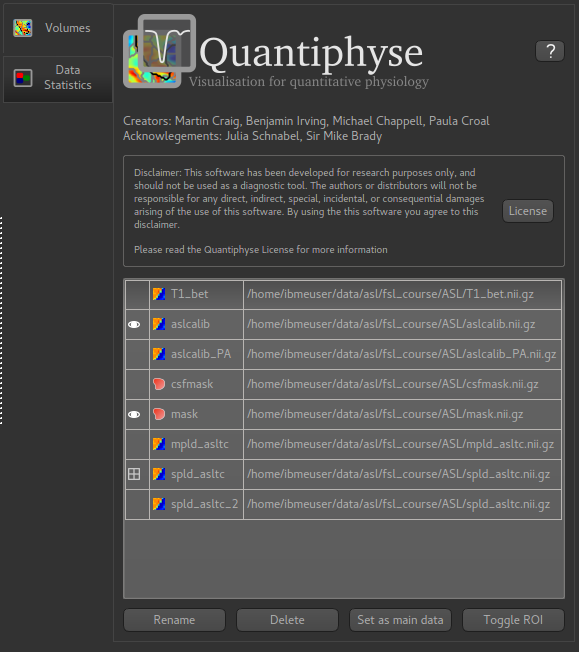
The Volumes List¶
After loading data it will appear in a list on the Volumes widget, which is always visible
by default:
The icon on the left indicates whether the data is visible or not:  indicates that this
is the main data (and will appear as a greyscale background),
indicates that this
is the main data (and will appear as a greyscale background),  indicates that this
data item is visible, either as an ROI or an overlay on top of the background.
indicates that this
data item is visible, either as an ROI or an overlay on top of the background.
Currently one overlay and one ROI are visible at a time. This limitation was intended to support the most common use case, but in the future the option to display multiple overlaid images will be added.
The icon next to the data name shows whether it is an  ROI or a
ROI or a  data set.
data set.
Using Widgets¶
Widgets appear to the right of the viewer window. Most widgets are accessed from the ‘Widgets’ menu above the viewer.
When selected, a widget will appear with a tab to the right of the viewer. You can switch between opened widgets by clicking on the tabs. A widget opened from the menu can be closed by clicking on the X in the top right of its tab.
Widgets may have very different user interfaces depending on what they do, however there are a number of common elements:
 Help button¶
Help button¶
This opens the online documentation page relevant to the widget. Internet access is required.
 Options button¶
Options button¶
This shows any extended options the widget may have. It is typically used by widgets which display plots as that limits the space available for options.
 Batch button¶
Batch button¶
This displays the batch code required to perform the widget’s processing, using the currently selected options. This can be useful when building batch files from interactive exploration. It is only supported by widgets which provide image processing functions.
 Citation¶
Citation¶
Many widgets are based around novel data processing techniques. The citation provides a reference to a published paper which can be used to find out more information about the underlying method. If you publish work using a widget with a citation, you should at the very least reference the paper given.

Clicking on the citation button performs an internet search for the paper.