 Quantiphyse¶
Quantiphyse¶
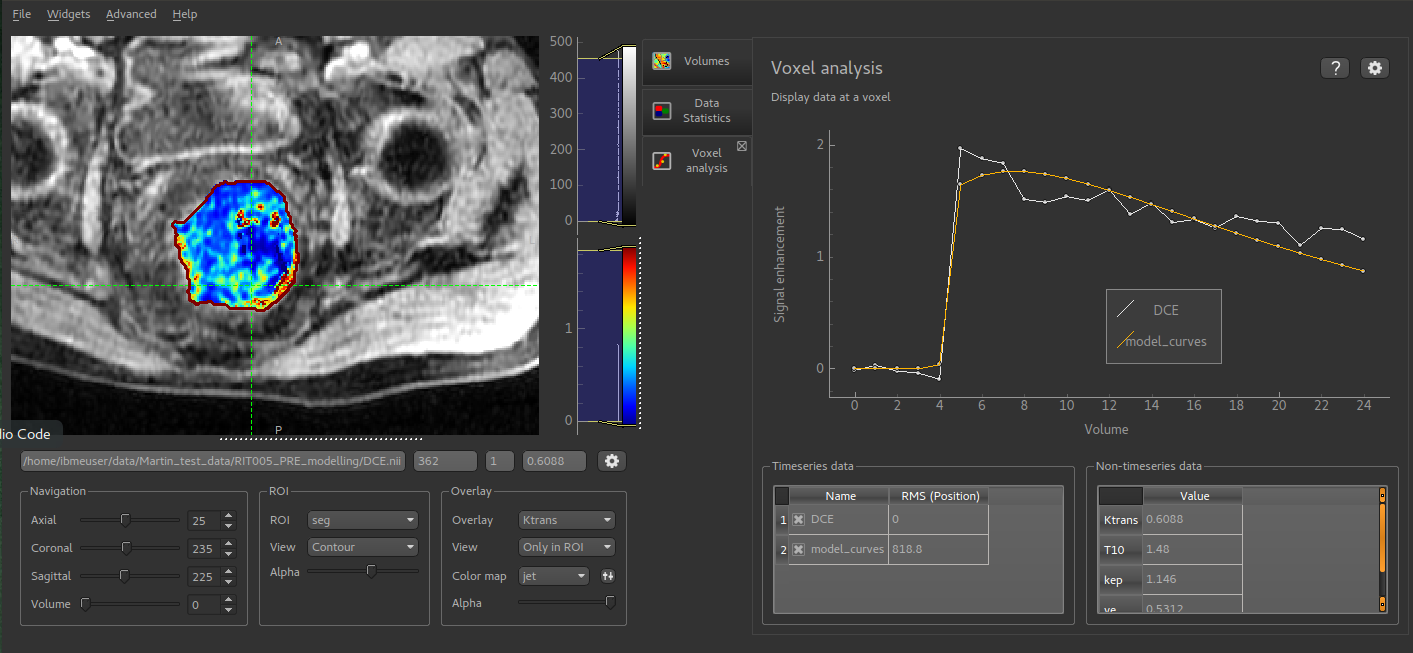
Quantiphyse is a visualisation and analysis tool for 3D and 4D biomedical data. It is particularly suited for physiological or functional imaging timeseries data.
Quantiphyse is built around the concept of making spatially resolved measurements of physical or physiological processes from imaging data using model-based or model-free methods, often exploiting Bayesian inference techniques.
Quantiphyse can analyse data by voxels or within regions of interest that may be manually or automatically created, e.g. using supervoxel or clustering methods.
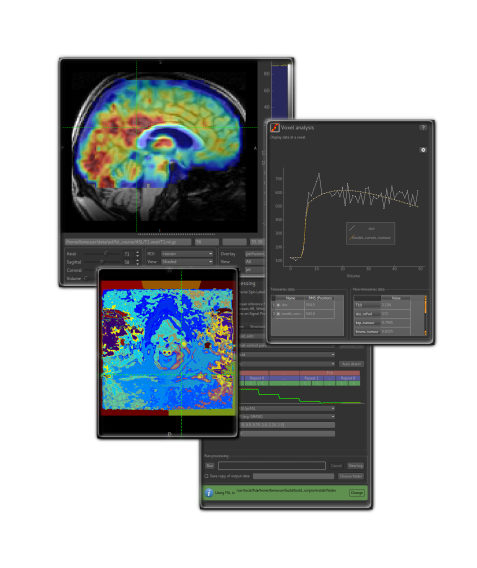
Features¶
- 2D orthographic visualisation and navigation of data, regions of interest (ROIs) and overlays
- Generic analysis tools including clustering, supervoxel generation and curve comparison
- Tools for CEST, ASL, DCE, DSC and qBOLD MRI analysis and modelling
- Integration with selected FSL tools
- ROI generation
- Registration and motion correction
- Extensible - see Quantiphyse plugins.
License¶
© 2017-2020 University of Oxford
Quantiphyse is Open Source software, licensed under the Apache Public License version 2.0
License details are displayed on first use and in the LICENSE file included in the distribution.
Note that this does not apply to all available plugins - you should check the licensing
terms for a plugin before using it.
Getting Quantiphyse¶
Quantiphyse is available on PyPi - see Installation of Quantiphyse.
Major releases of Quantiphyse are also available via the Oxford University Innovation Software Store. The packages held by OUI have no external dependencies and can be installed on Windows, Mac and Linux. They may lag behind the current PyPi release in terms of functionality.
User Guide¶
Bugs/Issues¶
Bugs may be submitted using the Github issue tracker for Quantiphyse.
For any other comments or feature requests please contact the current maintainer:
Contributors¶
- Martin Craig (Current developer)
- Ben Irving (‘Author of ‘PKview’, the predecessor to Quantiphyse)
- Michael Chappell
- Paula Croal
- Moss Zhao